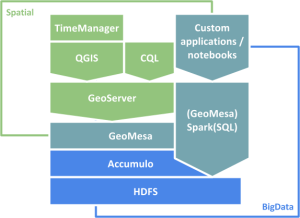
In a previous post, I showed how to use docker to run a single application (GeoServer) in a container and connect to it from your local QGIS install. Today’s post is about running a whole bunch of containers that interact with each other. More specifically, I’m using the images provided by Geodocker. The Geodocker repository provides a setup containing Accumulo, GeoMesa, and GeoServer. If you are not familiar with GeoMesa yet:
GeoMesa is an open-source, distributed, spatio-temporal database built on a number of distributed cloud data storage systems … GeoMesa aims to provide as much of the spatial querying and data manipulation to Accumulo as PostGIS does to Postgres.
The following sections show how to load data into GeoMesa, perform basic queries via command line, and finally publish data to GeoServer. The content is based largely on two GeoMesa tutorials: Geodocker: Bootstrapping GeoMesa Accumulo and Spark on AWS and Map-Reduce Ingest of GDELT, as well as Diethard Steiner’s post on Accumulo basics. The key difference is that this tutorial is written to be run locally (rather than on AWS or similar infrastructure) and that it spells out all user names and passwords preconfigured in Geodocker.
This guide was tested on Ubuntu and assumes that Docker is already installed. If you haven’t yet, you can install Docker as described in Install using the repository.
To get Geodocker set up, we need to get the code from Github and run the docker-compose command:
$ git clone https://github.com/geodocker/geodocker-geomesa.git
$ cd geodocker-geomesa/geodocker-accumulo-geomesa/
$ docker-compose up
This will take a while.
When docker-compose is finished, use a second console to check the status of all containers:
$ docker ps
CONTAINER ID IMAGE COMMAND CREATED STATUS PORTS NAMES
4a238494e15f quay.io/geomesa/accumulo-geomesa:latest "/sbin/entrypoint...." 19 hours ago Up 23 seconds geodockeraccumulogeomesa_accumulo-tserver_1
e2e0df3cae98 quay.io/geomesa/accumulo-geomesa:latest "/sbin/entrypoint...." 19 hours ago Up 22 seconds 0.0.0.0:50095->50095/tcp geodockeraccumulogeomesa_accumulo-monitor_1
e7056f552ef0 quay.io/geomesa/accumulo-geomesa:latest "/sbin/entrypoint...." 19 hours ago Up 24 seconds geodockeraccumulogeomesa_accumulo-master_1
dbc0ffa6c39c quay.io/geomesa/hdfs:latest "/sbin/entrypoint...." 19 hours ago Up 23 seconds geodockeraccumulogeomesa_hdfs-data_1
20e90a847c5b quay.io/geomesa/zookeeper:latest "/sbin/entrypoint...." 19 hours ago Up 24 seconds 2888/tcp, 0.0.0.0:2181->2181/tcp, 3888/tcp geodockeraccumulogeomesa_zookeeper_1
997b0e5d6699 quay.io/geomesa/geoserver:latest "/opt/tomcat/bin/c..." 19 hours ago Up 22 seconds 0.0.0.0:9090->9090/tcp geodockeraccumulogeomesa_geoserver_1
c17e149cda50 quay.io/geomesa/hdfs:latest "/sbin/entrypoint...." 19 hours ago Up 23 seconds 0.0.0.0:50070->50070/tcp geodockeraccumulogeomesa_hdfs-name_1
At the time of writing this post, the Geomesa version installed in this way is 1.3.2:
$ docker exec geodockeraccumulogeomesa_accumulo-master_1 geomesa version
GeoMesa tools version: 1.3.2
Commit ID: 2b66489e3d1dbe9464a9860925cca745198c637c
Branch: 2b66489e3d1dbe9464a9860925cca745198c637c
Build date: 2017-07-21T19:56:41+0000
Loading data
First we need to get some data. The available tutorials often refer to data published by the GDELT project. Let’s download data for three days, unzip it and copy it to the geodockeraccumulogeomesa_accumulo-master_1 container for further processing:
$ wget http://data.gdeltproject.org/events/20170710.export.CSV.zip
$ wget http://data.gdeltproject.org/events/20170711.export.CSV.zip
$ wget http://data.gdeltproject.org/events/20170712.export.CSV.zip
$ unzip 20170710.export.CSV.zip
$ unzip 20170711.export.CSV.zip
$ unzip 20170712.export.CSV.zip
$ docker cp ~/Downloads/geomesa/gdelt/20170710.export.CSV geodockeraccumulogeomesa_accumulo-master_1:/tmp/20170710.export.CSV
$ docker cp ~/Downloads/geomesa/gdelt/20170711.export.CSV geodockeraccumulogeomesa_accumulo-master_1:/tmp/20170711.export.CSV
$ docker cp ~/Downloads/geomesa/gdelt/20170712.export.CSV geodockeraccumulogeomesa_accumulo-master_1:/tmp/20170712.export.CSV
Loading or importing data is called “ingesting” in Geomesa parlance. Since the format of GDELT data is already predefined (the CSV mapping is defined in geomesa-tools/conf/sfts/gdelt/reference.conf), we can ingest the data:
$ docker exec geodockeraccumulogeomesa_accumulo-master_1 geomesa ingest -c geomesa.gdelt -C gdelt -f gdelt -s gdelt -u root -p GisPwd /tmp/20170710.export.CSV
$ docker exec geodockeraccumulogeomesa_accumulo-master_1 geomesa ingest -c geomesa.gdelt -C gdelt -f gdelt -s gdelt -u root -p GisPwd /tmp/20170711.export.CSV
$ docker exec geodockeraccumulogeomesa_accumulo-master_1 geomesa ingest -c geomesa.gdelt -C gdelt -f gdelt -s gdelt -u root -p GisPwd /tmp/20170712.export.CSV
Once the data is ingested, we can have a look at the the created table by asking GeoMesa to describe the created schema:
$ docker exec geodockeraccumulogeomesa_accumulo-master_1 geomesa describe-schema -c geomesa.gdelt -f gdelt -u root -p GisPwd
INFO Describing attributes of feature 'gdelt'
globalEventId | String
eventCode | String
eventBaseCode | String
eventRootCode | String
isRootEvent | Integer
actor1Name | String
actor1Code | String
actor1CountryCode | String
actor1GroupCode | String
actor1EthnicCode | String
actor1Religion1Code | String
actor1Religion2Code | String
actor2Name | String
actor2Code | String
actor2CountryCode | String
actor2GroupCode | String
actor2EthnicCode | String
actor2Religion1Code | String
actor2Religion2Code | String
quadClass | Integer
goldsteinScale | Double
numMentions | Integer
numSources | Integer
numArticles | Integer
avgTone | Double
dtg | Date (Spatio-temporally indexed)
geom | Point (Spatially indexed)
User data:
geomesa.index.dtg | dtg
geomesa.indices | z3:4:3,z2:3:3,records:2:3
geomesa.table.sharing | false
In the background, our data is stored in Accumulo tables. For a closer look, open an interactive terminal in the Accumulo master image:
$ docker exec -i -t geodockeraccumulogeomesa_accumulo-master_1 /bin/bash
and open the Accumulo shell:
# accumulo shell -u root -p GisPwd
When we store data in GeoMesa, there is not only one table but several. Each table has a specific purpose: storing metadata, records, or indexes. All tables get prefixed with the catalog table name:
root@accumulo> tables
accumulo.metadata
accumulo.replication
accumulo.root
geomesa.gdelt
geomesa.gdelt_gdelt_records_v2
geomesa.gdelt_gdelt_z2_v3
geomesa.gdelt_gdelt_z3_v4
geomesa.gdelt_queries
geomesa.gdelt_stats
By default, GeoMesa creates three indices:
Z2: for queries with a spatial component but no temporal component.
Z3: for queries with both a spatial and temporal component.
Record: for queries by feature ID.
But let’s get back to GeoMesa …
Querying data
Now we are ready to query the data. Let’s perform a simple attribute query first. Make sure that you are in the interactive terminal in the Accumulo master image:
$ docker exec -i -t geodockeraccumulogeomesa_accumulo-master_1 /bin/bash
This query filters for a certain event id:
# geomesa export -c geomesa.gdelt -f gdelt -u root -p GisPwd -q "globalEventId='671867776'"
Using GEOMESA_ACCUMULO_HOME = /opt/geomesa
id,globalEventId:String,eventCode:String,eventBaseCode:String,eventRootCode:String,isRootEvent:Integer,actor1Name:String,actor1Code:String,actor1CountryCode:String,actor1GroupCode:String,actor1EthnicCode:String,actor1Religion1Code:String,actor1Religion2Code:String,actor2Name:String,actor2Code:String,actor2CountryCode:String,actor2GroupCode:String,actor2EthnicCode:String,actor2Religion1Code:String,actor2Religion2Code:String,quadClass:Integer,goldsteinScale:Double,numMentions:Integer,numSources:Integer,numArticles:Integer,avgTone:Double,dtg:Date,*geom:Point:srid=4326
d9e6ab555785827f4e5f03d6810bbf05,671867776,120,120,12,1,UNITED STATES,USA,USA,,,,,,,,,,,,3,-4.0,20,2,20,8.77192982456137,2007-07-13T00:00:00.000Z,POINT (-97 38)
INFO Feature export complete to standard out in 2290ms for 1 features
If the attribute query runs successfully, we can advance to some geo goodness … that’s why we are interested in GeoMesa after all … and perform a spatial query:
# geomesa export -c geomesa.gdelt -f gdelt -u root -p GisPwd -q "CONTAINS(POLYGON ((0 0, 0 90, 90 90, 90 0, 0 0)),geom)" -m 3
Using GEOMESA_ACCUMULO_HOME = /opt/geomesa
id,globalEventId:String,eventCode:String,eventBaseCode:String,eventRootCode:String,isRootEvent:Integer,actor1Name:String,actor1Code:String,actor1CountryCode:String,actor1GroupCode:String,actor1EthnicCode:String,actor1Religion1Code:String,actor1Religion2Code:String,actor2Name:String,actor2Code:String,actor2CountryCode:String,actor2GroupCode:String,actor2EthnicCode:String,actor2Religion1Code:String,actor2Religion2Code:String,quadClass:Integer,goldsteinScale:Double,numMentions:Integer,numSources:Integer,numArticles:Integer,avgTone:Double,dtg:Date,*geom:Point:srid=4326
139346754923c07e4f6a3ee01a3f7d83,671713129,030,030,03,1,NIGERIA,NGA,NGA,,,,,LIBYA,LBY,LBY,,,,,1,4.0,16,2,16,-1.4060533085217,2017-07-10T00:00:00.000Z,POINT (5.43827 5.35886)
9e8e885e63116253956e40132c62c139,671928676,042,042,04,1,NIGERIA,NGA,NGA,,,,,OPEC,IGOBUSOPC,,OPC,,,,1,1.9,5,1,5,-0.90909090909091,2017-07-10T00:00:00.000Z,POINT (5.43827 5.35886)
d6c6162d83c72bc369f68bcb4b992e2d,671817380,043,043,04,0,OPEC,IGOBUSOPC,,OPC,,,,RUSSIA,RUS,RUS,,,,,1,2.8,2,1,2,-1.59453302961275,2017-07-09T00:00:00.000Z,POINT (5.43827 5.35886)
INFO Feature export complete to standard out in 2127ms for 3 features
Functions that can be used in export command queries/filters are (E)CQL functions from geotools for the most part. More sophisticated queries require SparkSQL.
Publishing GeoMesa tables with GeoServer
To view data in GeoServer, go to http://localhost:9090/geoserver/web. Login with admin:geoserver.
First, we create a new workspace called “geomesa”.
Then, we can create a new store of type Accumulo (GeoMesa) called “gdelt”. Use the following parameters:
instanceId = accumulo
zookeepers = zookeeper
user = root
password = GisPwd
tableName = geomesa.gdelt

Then we can configure a Layer that publishes the content of our new data store. It is good to check the coordinate reference system settings and insert the bounding box information:
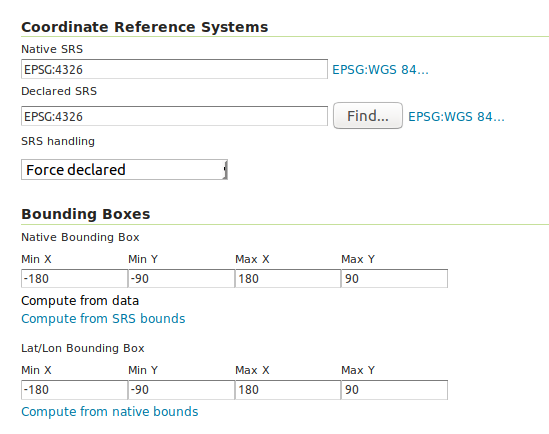
To preview the WMS, go to GeoServer’s preview:
http://localhost:9090/geoserver/geomesa/wms?service=WMS&version=1.1.0&request=GetMap&layers=geomesa:gdelt&styles=&bbox=-180.0,-90.0,180.0,90.0&width=768&height=384&srs=EPSG:4326&format=application/openlayers&TIME=2017-07-10T00:00:00.000Z/2017-07-10T01:00:00.000Z#
Which will look something like this:

GeoMesa data filtered using CQL in GeoServer preview
For more display options, check the official GeoMesa tutorial.
If you check the preview URL more closely, you will notice that it specifies a time window:
&TIME=2017-07-10T00:00:00.000Z/2017-07-10T01:00:00.000Z
This is exactly where QGIS TimeManager could come in: Using TimeManager for WMS-T layers. Interoperatbility for the win!